Contamination Controls for Analysis of Root Canal Samples by Molecular Methods: An Overlooked and Unsolved Problem
Abstract
Introduction: It has been almost 20 years since molec- ular methods were first described for the analysis of root canal microbial flora. Contamination control samples are essential to establish DNA decontamination before taking root canal samples, and this review assessed those studies. Methods: Using PubMed, a search was conducted for studies using molecular microbial analysis for the investigation of endodontic samples. Studies were grouped according to the cleaning protocol, acqui- sition methods, and processing of control samples taken to check for contamination. Results: Of 136 studies applying molecular analysis to root canal samples, 21 studies performed surface cleaning and checking nucle- otide decontamination with contamination control sam- ples processed by polymerase chain reaction. Only 1 study described disinfection, sampling from the access cavity, and processing by polymerase chain reaction and reported the result; that study reported that all samples contained contaminating bacterial DNA. Conclusions: Cleaning, disinfection, and checking for contamination are basic scientific prerequisites for this type of investigation; yet, this review identifies it as an overlooked issue. On the basis of this review, we call for improved scientific practice in this field. (J Endod 2016;42:1003–1008)
Key Words
Contamination controls, molecular analysis, polymerase chain reaction, root canal sampling.
The first published article applying molecular anal- ysis for the investigation of root canal microbiota was published in 1997 when the polymerase chain reac- tion (PCR) technique was used to target specific DNA sequences in samples taken from infected root canals(1). Advances in molecular techniques for microbial identification provide fast, accurate, and straightfor- ward procedures for taxo-nomic characterization of species in clinical samples. From that first publication in 1997 to October 2015, there have been more than 136 articles that use a molecular approach for laboratory processing of the microbial isolates. In this narrative review, we examine an important specific issue, namely contamination controls for the analysis of root canal samples using molecular methods.
A Brief History
Before the advent of sequence-based identification, microbiological analysis of root canal microbiota relied on conventional culture. The principles and scientific methods for microbial sampling from infected root canals were developed in the 1960s and published in a classic thesis (2). Moller’s work was significant for several reasons, not the least being the scientific rigor he applied to controlling the field before taking a sample from the subject area. Numerous articles using culture techniques for processing root canal samples had been published before that time. What changed was the logical yet simple refinements made by Moller to rigorously control the sampling and subsequent laboratory processing for the accurate recovery of bacteria.
An essential preparatory step was to work in a sterile field that excluded extra- neous microbes and then check the field with a contamination control sample to confirm the absence of contaminants before entry into the pulp space. This seemingly simple but hitherto overlooked step was based on the premise that if microbes were present on the tooth surface, they may be picked up when taking the sample, thus contaminating the results of the root canal sample. Moller asked 2 essential ques- tions: How does one clean and disinfect the tooth surface? and, when taking the sam-ple, How can one check that the operating field is free from surface-contaminating microbes? M€oller answered those questions by exhaustively developing and testing various cleaning and disinfection protocols that could be applied in a reproducible way. He then confirmed the validity of his methods by taking contamination controls to check the effectiveness of cleaning and disinfection steps. Before his careful pio- neering work, many studies were poorly controlled for contamination, and there were many gaping technical problems that negated accurate sampling of the infected root canal space.
What You Can’t See Won’t Matter or Will It?
It is important to recognize the essential differences between cul- ture and molecular-based studies. Conventional culture reveals the presence of living, growing microbes that can be seen as colonies on agar plates or broth culture. Cultivated colonies can then be processed by phenotypic and biochemical tests to identify the species. Molecular studies record the presence of DNA, usually specific nucleotide se- quences targeted by PCR. In recent years, advances in DNA sequencing mean that recovered DNA can be partially or wholly sequenced for spe- cies identification. Because these are such disparate methods, it is worth noting that sterility means something different for each method. A sterile field for culture means that there are no cultivable viable microbes, whereas a sterile field for molecular work means an absence of recover- able microbial nucleotides from the respective sampled region.
The relevant questions posed and effectively answered by M€oller in his 1966 work are as pertinent today as they were in the 1960s. Is the tooth surface free from contaminating microbial DNA before taking the root canal sample? The corollary question is can it be confirmed that cleaning and decontamination (of microbial nucleotides) have been effective? Have contamination controls been taken to ensure that extraneous DNA is not picked up when taking a sample from the root canal space?
A Challenging Issue: DNA Binding to Hydroxyapatite
The chemical composition of teeth presents a special issue in relation to nucleotide sampling because of the strong binding affinity of hydroxy- apatite for DNA. This discovery (3) provided great benefit for the separa- tion of DNA in early studies (4). It provides a unique scientific advantage, such as the ability to recover ancient DNA from animal and human remains
(5). Yet, it also presents special challenges in endodontic microbiology. Recent studies have confirmed both the binding affinity of hydroxy- apatite and dentin for DNA (6, 7). One of the implications from this work is that microbial biofilms, which contain extracellular DNA (8), potentially leave long-lasting traces of DNA highly bound to the tooth surface. Thus, any microbial biofilm that has had intimate contact with any part of the tooth has the potential to leave detectable traces of microbial DNA for extended periods. Reliable removal of the bound
DNA presents an as-yet unsolved challenge.
In this review, we assessed studies that have used molecular approaches for the analysis of root canal microbiota and evaluated their cleaning, disinfection, and nucleotide decontamination protocol; whether a contamination control sample was taken; and how it was pro- cessed in the laboratory. Finally, we critically evaluated the results of the contamination controls.
Materials and Methods
Using PubMed, a search was conducted for the following terms: ‘‘PCR + root canal,’’ ‘‘PCR + endodontic,’’ ‘‘molecular + endodontic,’’ and ‘‘molecular + root canal.’’ Of those articles describing in vivo inves- tigation of root canal microbes, 136 were selected as fulfilling the search criteria and suitable for analysis. The articles were grouped according to whether the surface was cleaned, if it was checked with a contamination control sample, and how samples were processed in the laboratory and then divided into categories. Studies that undertook surface cleaning, acquired a contamination control sample, and processed it by PCR were selected for detailed analysis.
Results and Discussion
There were 136 articles that fulfilled the search criteria. These studies were separated initially according to whether a contamination control sample was taken and how the control sample was processed in the laboratory (Fig. 1). Eight of 136 articles used no or had no infor- mation about a surface decontamination procedure (Fig. 1). The appli- cation of appropriate controls is a fundamental requirement in any scientific inquiry, so without controls it calls into question how those articles reached scientific publication.
The majority of articles (89/136) reported cleaning of the tooth surface but no description of a contamination control sample (Fig. 1). An obvious limitation of cleaning without taking a control sam- ple is that there are no data to support the contention that the root canal samples actually disclose microbes exclusively located within the root canal space.
In 16 of 136 articles, the tooth surface was cleaned, and the ste- rility was checked by culture (Fig. 1). Providing that the results were negative (ie, no samples were positive by culture), this means that there were no living, cultivable bacteria, which is a positive step to confirm a sterile field. However, a control sample processed by culture is insuffi- cient for molecular work because the molecular approach targets nucleotide sequences. For the study to be effective and provide authentic sampling results, the operating field must be tested for the presence of microbial DNA. There should be no amplifiable DNA, which helps verify that the sequences isolated are validly derived from the root canal space. Only 23 of 136 articles were identified that performed surface cleaning and checking nucleotide decontamination by taking a contam- ination control sample processed by PCR (Fig. 1). These 23 articles were evaluated according to their reporting of contamination controls (Fig. 2). Of these, 2 articles (9, 10) used the same material from earlier studies (11, 12) (Fig. 2), so they were not analyzed further. In 3 of 23 articles (11, 13, 14), no results were reported from contamination controls, which leaves the findings unknown. It would have been a sound scientific procedure for these articles to report the results of such controls.
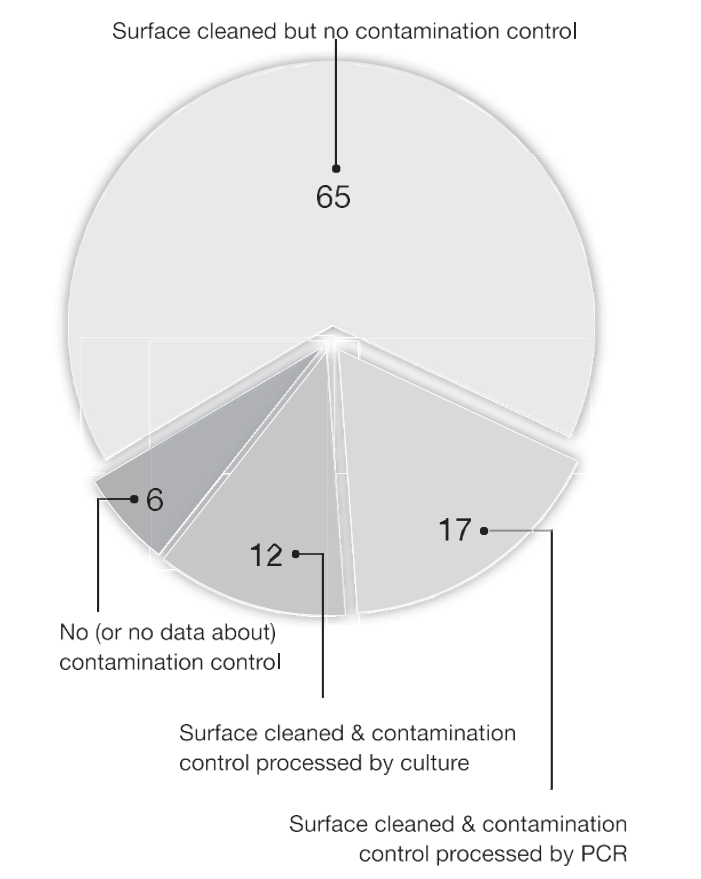
Figure 1. A pie chart showing analysis of 136 articles separated by whether a contamination control sample was taken and whether the control sample was laboratory processed by culture or PCR. The numbers are the percent of articles out of total 136 articles.
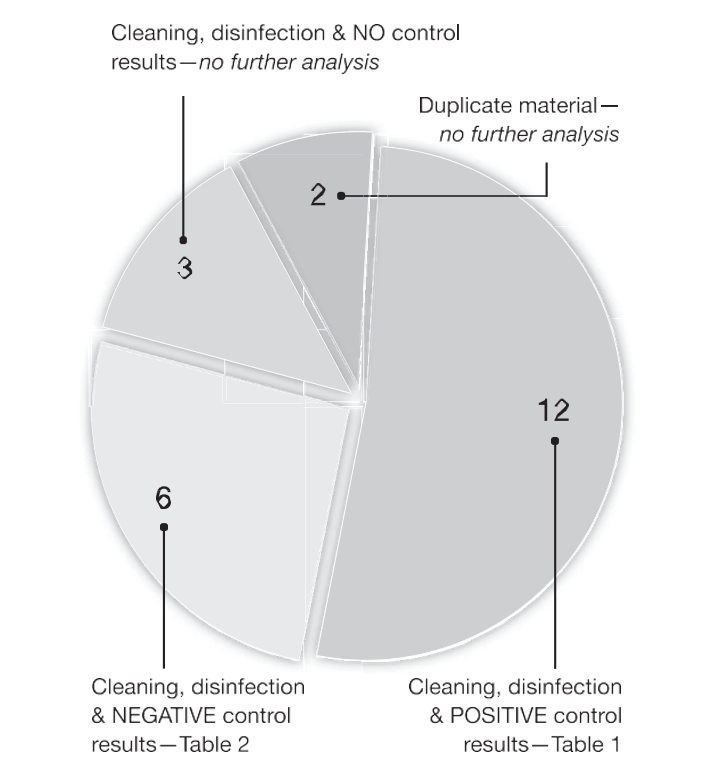
Figure 2. A pie chart showing analysis of 23 articles that performed surface cleaning and checking nucleotide decontamination by taking a contamination control sample processed by PCR. Figures shown are the number of articles in each group (from a total of 23 articles) and how 18 articles are assigned to Tables 1 and 2.
Thus, there was no further analysis of these 3 studies (Fig. 2). From these 23 articles, there were 18 that performed surface cleaning, checked with a contamination control sample processed by PCR, and reported the results. These 18 papers were further divided into 2 groups (Fig. 2): 12 articles with some contamination control re- sults that were positive for residual DNA (Table 1) and 6 articles with all- negative control results (Table 2).
Of the 12 articles with positive results for DNA contamination, 10 described exclusion of those teeth from the case material (Table 1). One article (16) described all contamination control samples as posi- tive for bacterial DNA, which might negate the reported findings from the root canal samples; however, the species selected for investigation in that study, Enterococcus faecalis, was not detected in species specific contamination controls, so the authors proceeded with analysis of the root canal samples (16). In another article (15), control samples were taken in only 7 of 58 teeth, and 1 case with a positive contamina- tion control result was not excluded from the study (Table 1).
In 6 of 18 articles that included the contamination control results, all contamination control samples were reported as negative (Table 2). This finding stands in stark contrast to the majority of other work that consistently indicates great difficulty in obtaining nucleotide-free tooth surfaces. The specific reasons are unknown but may include PCR primer sensitivity, the number of amplification cycles (too low may fail to amplify DNA), the method of DNA extraction (eg, boiling cells, used in 1 study, may not release DNA from some species), and whether the primers were species specific or universal (for recovery of all bacterial DNA). How and where the control sample was taken may also significantly influence the result. Some of these issues are further addressed later.
TABLE 1. Analysis of 12 Articles that Describe Cleaning, Disinfection, and a Positive Surface Contamination Control Result by Polymerase Chain Reaction
|
Study |
Method of disinfection* |
Site of control sample | Primer type used for analysis | Control results (positive/total) |
| Kaufman et al, 2005 (15) | 30% H2O2, 5% I2, 6% NaOCl | Surface | Enterococcal specific | 1/7† |
| Sedgley et al, 2006 (16) | 30% H2O2, 6% NaOCl | Cavity | Universal and
Enterococcal specific |
23/23‡ |
| Ro^c¸ as et al, 2011 (17) | 3% H2O2, 2.5% NaOCl | Surface | Universal | 3/50 |
| Ro^c¸ as et al, 2011 (18) | 3% H2O2, 2.5% NaOCl | Surface | Universal | 3/27 |
| Paiva et al, 2012 (12) | 6% H2O2, 2% I2, 6% H2O2, 2.5% NaOCl | Cavosurface angle | Universal | 3/30 |
| Ro^c¸ as et al, 2012 (19) | 3% H2O2, 2.5% NaOCl | Operative field | Universal | 8/50 |
| Neves et al, 2013 (20) | 6% H2O2, 2.5% NaOCl | Cavosurface angle | Universal | 5/50 |
| Paiva et al, 2013 (21) | 6% H2O2, 2% I2, 6% H2O2, 2.5% NaOCl | Cavosurface angle | Universal | 4/18 |
| Ro^c¸ as et al, 2013 (22) | 3% H2O2, 2.5% NaOCl | Surface | Universal | 1/50 |
| Ro^c¸ as et al, 2013 (23) | 6% H2O2, 2% I2, 6% H2O2, 2.5% NaOCl | Cavosurface angle | Universal | 5/45 |
| Ro^c¸ as et al, 2014 (24) | 6% H2O2, 2.5% NaOCl | Cavosurface angle | Universal | 3/28 |
| Pinheiro et al, 2015 (25) | 30% H2O2, 2.5% NaOCl | Surface | Enterococcal specific | 2/20 |
H2O2, hydrogen peroxide; I, iodine; NaOCl, sodium hypochlorite.
*Two cleans described in all articles but not stated in 1 study (16).
†Controls done in 7 cases only (from a total 58); 1 case with a positive contamination control result was not excluded from the study.
‡23/23 for Universal; 0/23 for Enterococcal primers. All cases were included in the study material.
TABLE 2. Analysis of 6 Articles that Describe Cleaning, Disinfection, and Negative Surface Contamination Control Results by Polymerase Chain Reaction
|
Study |
Method of disinfection |
No. of cleans |
Site of control sample |
Primer type used for analysis | Processed controls (n/total) |
| Tomazhino et al, 2007 (26) | 30% H2O2, 5% I2 | 1 | Surface | BPR specific | 100/100 |
| Montagner et al, 2012 (27) | 30% H2O2, 2.5% NaOCl | 1 | Surface | Universal | 20/20 |
| Gomes et al, 2013 (28) | 30% H2O2, 2.5% NaOCl | 1 | Operative field | Universal | 15/15 |
| Stojanovi´c et al, 2014 (29) | 30% H2O2, 2.5% NaOCl | 2 | Operative field | Universal | 51/51 |
| Teles et al, 2014 (30) | 3% H2O2, 3% NaOCl | 2 | Operative field | NS | 69/69 |
| Moraes et al, 2015 (31) | 30% H2O2, 5.25% NaOCl | 1 | Operative field and surface | Universal | 27/27 |
BPR, black-pigmented rods; H2O2, hydrogen peroxide; I2, iodine; NaOCl, sodium hypochlorite; NS, not stated.
Are All Cleaning Protocols Effective?
Another important question is what were the cleaning and disin- fection protocols applied to the tooth surface before taking a sample? For culture-based studies, M€oller showed that cleaning with hydrogen peroxide (H2O2) and disinfection with tincture of iodine were highly effective for producing a sterile tooth surface. The iodine was inactivated with sodium thiosulfate (to prevent carryover of iodine into culture media) before taking a contamination control sample. Does the same protocol apply for decontamination before taking a control sample for molecular analysis?
The only study known to the authors conducted on cleaning pro- tocols for molecular sampling (32) showed that after cleaning with H2O2, the use of iodine for disinfection resulted in almost half the teeth still being contaminated with recoverable DNA. An improvement occurred when sodium hypochlorite (NaOCl) was used for decontam- ination, resulting in DNA detection in 13% of samples compared with 45% after using iodine (32). Interestingly, 1 article described the use of iodine for disinfection (Table 2) yet reported no positive findings out of 100 cases (26), which stands in contrast to the findings described in the cleaning protocols study (32).
The fallibility of current decontamination methods for molecular work is also shown in a study (16) that used H2O2 and 6% NaOCl to clean and disinfect the tooth surfaces of 23 teeth before sampling the root canal for E. faecalis. After cleaning, contamination control sam- ples taken from the access cavity and processed with universal primers (for all bacterial DNA) revealed recoverable DNA in 100% of samples (16), which led the authors to raise the question as to whether total physical removal of DNA remaining after disinfecting endodontic access cavities might ever be achievable.
Of 136 articles using molecular methods for analysis of root canal samples, there was a nonuniform approach used for surface cleaning and disinfection. Most studies used a combination of H2O2 and NaOCl for cleaning and disinfection of the operative field and tooth (Fig. 3). However, 16% of articles (Fig. 3) used cleaning protocols that did not describe the use of NaOCl, which is generally regarded as essential for the removal of DNA.
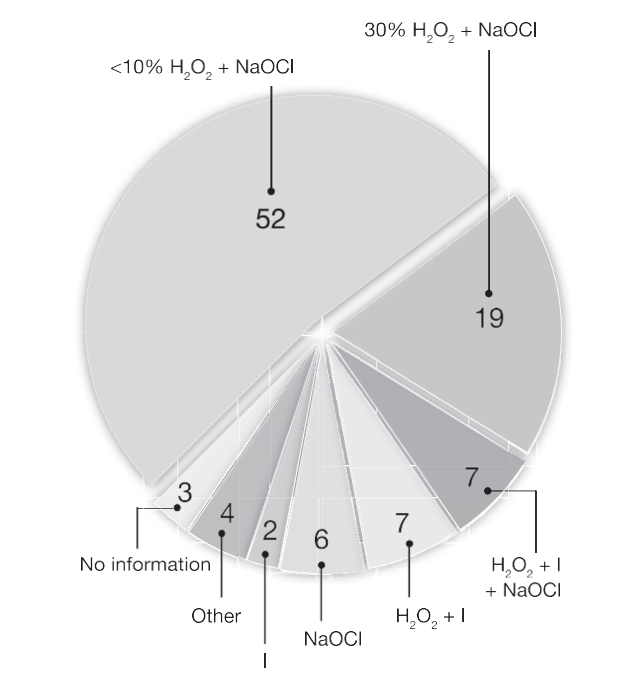
Figure 3. A pie chart showing analysis of 136 articles based on the cleaning and disinfection protocol. The numbers are the percent of articles out of total 136 articles.
When and Where Should a Contamination Control Sample Be Taken?
In the oral cavity, microbial DNA is everywhere. When a sample is taken for the purpose of molecular microbial analysis of the root canal space, the aim should be to first exclude all extraneous DNA and then to verify its exclusion by taking a contamination control sample. That sam- ple must be processed using molecular methods to ascertain the absence of any residual microbial DNA.
When should the sample be taken? If it is taken after cleaning the tooth surface but before access cavity preparation, this would test for residual DNA before cutting the access cavity. Yet, it is likely that preparation of the access cavity will lead to further microbial DNA deposition or uncover existing DNA within the entry site. Therefore, after access cavity preparation, a second cleaning and disinfection should be performed before taking a sample (33). This is the reason for the recommendation of a 2-step disinfection protocol (32). Briefly, the tooth and surrounding field are cleansed with 30% H2O2 and decontaminated with a 2.5% (or higher concentration) NaOCl solution. After access cavity preparation using sterile burs without water spray, the operative field, including the pulp chamber, is then swabbed again with 2.5% NaOCl. NaOCl is then inactivated by sterile 5% sodium thiosulfate (2, 32).
It is interesting to note that a 2-step clean was performed in 11 of 12 studies that described some positive results for contamination con- trol samples (Table 1); yet, 4 of 6 studies that reported negative results for their control (Table 2) used a single-clean protocol.
Where should the sample be taken? It is expected that the operative field (rubber dam and retainer) should be sterile after decontamination and disinfection. In 4 of 6 studies that had negative contamination con- trol results (Table 2), samples were taken from the operative field so it is not surprising that these sites were free of contamination. It is also reasonable to expect that the tooth surface should have no recoverable microbial DNA before taking a root canal sample.
Of the 18 articles in Tables 1 and 2, 13 described samples taken at the tooth surface (6 of which described cavosurface angles, presumably at the surface side of the access cavity); 10 of 15 described some positive contamination control samples (Table 1), and 3 reported negative results (Table 2). However, the external tooth surface is not the site for taking the root canal sample.
Because the aim is to sample the root canal contents, the site of greatest relevance is the access cavity; it is reasonable to expect that the cavity walls are free of contaminating DNA before entry is made into the pulp chamber. Only 2 of the 136 studies identified in this review (11, 16) described 2-step cleaning, taking a contamination con- trol sample from the access cavity before entry into the pulp space and processing the sample with universal PCR primers targeted broadly for bacterial DNA. One of these (11) did not report the results, so the findings from these controls are unknown. The other (16) reported that 100% of access cavities tested were positive for contaminating bacterial DNA.
If a sample is taken after breaching the infected pulp chamber, there is either the risk that microbial DNA from the root canal will be picked up in the control sample or that disinfection of the access cavity before taking the control will ruin the subsequent root canal sample. From the foregoing analysis, it is apparent that it is difficult to elim- inate all traces of bacterial DNA even if the bacteria in the access cavity are dead. It is also evident that contamination control sampling has been mostly taken at sites that are less than relevant for taking a contamination control sample. The site of greatest importance is the access cavity—af- ter it is prepared and before breaching the pulp chamber. Only 1 of 136 studies has done so and reported the findings (16). On the basis of what is currently known, the optimal time and site are to take the control sam- ple of the dentin walls of the access cavity after a 2-step cleaning and disinfection and before entry into the root canal space.
Sensitivity of PCR
The PCR technique is widely regarded as highly sensitive for the recovery of DNA. There are many factors that influence the ability of PCR to detect nucleotide sequences, and these include primer design, amplicon size, inhibition factors, and cycle number (34–36). Higher cycle numbers increase sensitivity, and it is generally regarded that 35 to 40 cycles provide optimum amplification with limited artifacts. All but 1 of the 18 studies described in Tables 1 and 2 used S35 cycles. One study used 30-cycle amplification and reported no recoverable DNA from 69 contamination control samples (30).
How sensitive is PCR when applied to samples taken for contam- ination controls? One way to calibrate sensitivity is to measure the detec- tion limit (ie, the minimum number of copies/cells that can be detected by that method). Of the studies in Tables 1 and 2, 16 of 18 did not describe a detection limit. Two studies did so; both articles (17, 18) report a thousand cell detection limit.
Live or Dead?
The subject of whether the contaminating DNA is derived from live or dead cells is outside the scope of this review. However, it is worth noting briefly that although a molecular marker for DNA that delineates living from dead cells would be ideal, no such reliable technology is currently available. Various molecular approaches for the preferential detection of viable cells have been proposed, such as ethidium mono- azide and propidium monoazide (37, 38), but there are no unanimously agreed-upon molecular markers for living cells. Because messenger RNA (mRNA) is expressed in living cells and has a short half- life, the presence of measurable mRNA is one such putative molecular indicator of living microbes. However, there are few studies that have applied it to endodontic microbiology (39, 40), probably because of the many challenges that make it difficult to accurately detect and measure mRNA in clinical samples.
Plain Speaking on Conducting Suitable Molecular Decontamination Controls
The paucity of studies using satisfactory molecular decontamination controls raises the question as to whether it is possible to satisfactorily clean microbe-covered enamel, restorative materials, dentin, and the ac- cess cavity to achieve a nucleotide-free field. Is there an unstated reason why so many studies have not done or reported on the results of DNA decontamination controls? It seems so, and it ought to be brought out. As mentioned previously, only 1 of 136 studies identified in this review described current best practice cleaning, disinfection, and site- specific (ie, access cavity) contamination control samples and processed them with universal primers that detect all bacterial DNA (16). That study reported that 100% of contamination control samples were positive for contaminating microbial DNA (16). In our own preliminary investiga- tions, we found that cleaning the surface with 30% H2O2, 2.5% NaOCl, and 5% iodine followed by inactivation of NaOCl with thiosulfate is insuf- ficient to eliminate bacterial DNA from in vivo access cavities. Contam- ination control samples from the access cavity were positive for bacterial DNA in 35%–97% depending on primer choice (unpublished data).
Based on these reports and our own observations, it is time to bring the issue of contamination controls to the surface. Failure to satis- factorily clean and disinfect the tooth surface and access cavity and then checking it with a contamination control before taking a root canal sam- ple for molecular analysis is a problem that must not be overlooked. It is an issue that readers should be aware of when reading endodontic microbiology studies in scientific journals, and it is reasonable to expect that authors and journal referees ensure that all appropriate steps have been performed to a validated scientific standard. When this has not been done, it ought to be raised in the discussion section so that it is appropriately acknowledged and discussed in the article.
Where to from Here?
Fifty years after M€oller’s contribution, it is time for a bold re-examination of the controls applied in molecular analysis of sam- ples from the root canal space. Tooth structure, in particular the root canal space, has unique challenges for microbial sampling and molecular processing, so changes are warranted in the way we deal with this issue. Based on the foregoing discussion, it is impor- tant to review and reassess the problem. Those conducting a clinical study of this kind know the enormous amount of work to plan it, recruit patients, collect the clinical material, and undertake the lab- oratory work. It makes good sense to ensure that the data obtained are accurate, reliable, and true.
Now is the time to re-establish basic scientific principles. Valid sampling must begin with proper cleaning and nucleotide disinfection of the tooth surface. Proper protocols are needed for this purpose, and the sterility of the field must be checked by appropriate molecular contamination controls. The results should be clearly articulated in articles prepared for publication. Thus, for cleaning, disinfection, and contamination controls, we propose the following:
- Clinicians and researchers working in this field of inquiry reintro- duce the practice.
- Referees and editors should insist on
- Readers look for and place greater value on articles that adhere to and describe the
Conclusions
This review assesses the practice of disinfection in studies using molecular analysis of the root canal microbiota and the procedures applied for verifying nucleotide decontamination. Microbial DNA is ubiquitous on tooth surfaces, and the high binding affinity between DNA and hydroxyapatite poses a special challenge for decontamination. The lessons learned 50 years ago for culture methods (excluding and eliminating contamination and then checking for it) appear to have been largely overlooked in studies using molecular methods for the analysis of root canal microbiota. Scrutiny of these studies revealed that only 1 of 136 reported current best practice cleaning, disinfection, and taking a site-specific (access cavity) contamination control sample processed by PCR (16). A salient finding from that study was that all control samples taken from the access cavity were positive for contam- inating DNA (16), which highlights just how difficult the task is to achieve a fully nucleotide-decontaminated site. It is a basic scientific principle that the relevant site for sampling undergo thorough nucleo- tide disinfection and that there are contamination controls to ensure that it is effective. After almost 20 years and over 136 publications of molecular techniques for sampling the infected root canal, it is time for advanced methods to get back to basics.
Acknowledgments
The authors thank Drs G€oran Sundqvist and Ramachandran Nair for critical review of the manuscript.
The authors deny any conflicts of interest related to this study.